Two advanced tools for understanding antimicrobial resistance
EPI assays and qPCR are two of several tools that BIOMIN scientists and researchers use to better understand and potentially combat the mechanisms of antibiotic resistance.
The spread of antibiotic resistant bacteria pose a major threat to modern medicine (WHO, 2014). Extensive antibiotic use in agriculture is one of many factors that may contribute to antibiotic resistance (Ventola, 2015). A key issue is the development of multi-drug resistance (MDR) in pathogenic bacteria found in the digestive tract.
Globally, there is a continuing need to prevent the further increase in MDR bacteria, not just in clinical, but in natural environments, too. In order to come up with successful strategies to battle the spread of antimicrobial resistance in animals, certain feed additives with competitive properties (e.g. that inhibit the growth of pathogenic bacteria) might also disrupt resistance mechanisms.
At the BIOMIN Research Center, we use several advanced analytical methods to better understand and potentially combat the mechanisms of antibiotic resistance. First, in vitro efflux pump inhibitor assays allow us to identify substances as potential resistance inhibitors in the laboratory. Second, cutting-edge metagenomics technologies allow us to detect and track bacterial genes or elements in complex environments using samples sourced from farms.
How multi-drug resistance happens
Multi-drug resistance in bacteria occurs by the accumulation of resistance genes on resistance plasmids,with each gene coding for resistance to a specific agent (Figure 1A), and/or by the action of multidrug efflux pumps, which can pump out more than one antibiotic drug (Figure 1C).
Resistance plasmids are often transferred very efficiently from cell to cell (Figure 1B). Resistance by efflux pumps occurs by the increased expression of genes that code for these pumps. Some pumps in Gram-negative bacteria (e.g. AcrAB-TolC in Salmonella) are especially important because they can pump out most of the antibiotics currently in use.
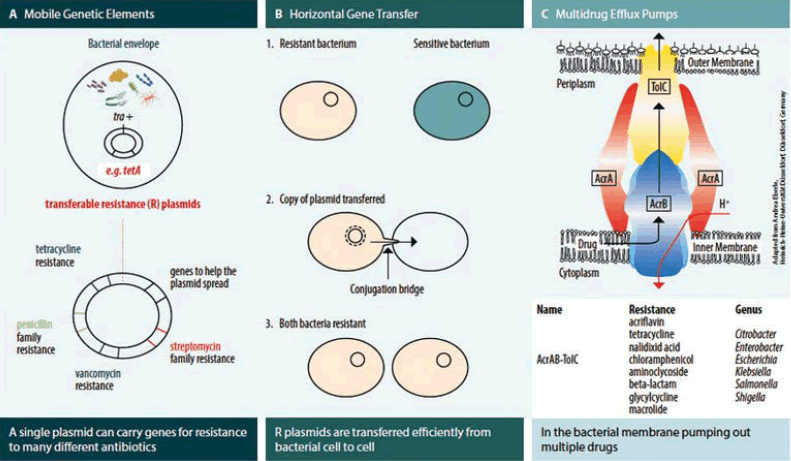
A) Multidrug resistant bacteria often carry mobile genetic elements like resistance plasmids, which can acquire many resistance genes through gene accumulation. B) Horizontal gene transfer via resistance plasmids efficiently passes resistance genes from one bacterium to another, contributing to the spread of antibiotic resistance in bacterial populations. C) Another mechanism of multidrug resistance is the active pumping out of drugs by multidrug efflux pumps.
Multidrug efflux: a key target in reversing antibiotic resistance
One way of prolonging antibiotic efficiency against multidrug-resistant pathogens is by blocking their efflux pumps with efflux pump inhibitors (EPIs). Natural plant-derived substances (phytogenics) have emerged as promising candidates, capable of improving the potency of antibiotics even at low concentrations, and preventing the emergence of resistance.
Efflux activity can be directly measured by fluorescence-based assays, based on two principles. First, various fluorescent dyes will shift in color and intensity when they enter the lipophilic environment inside of bacterial cells. Second, these dyes are actively pumped out of the cells by the efflux machinery. Monitoring the shifts in fluorescence enables us to see how fast the bacteria can pump out dyes, and, if an added substance is a potential inhibitor (Figure 2).
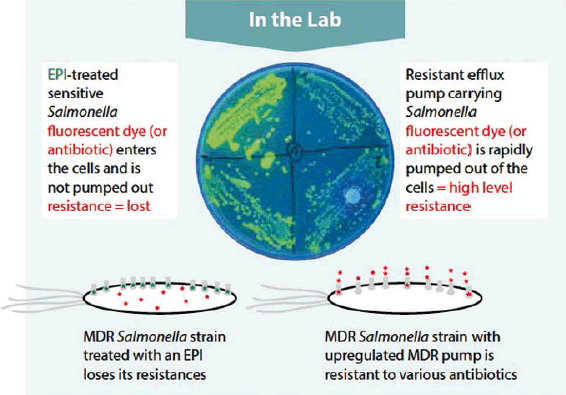
Efflux pump inhibitorassay results
In experiments at the BIOMIN Research Center, a Salmonella enterica serovar Typhimurium strain carrying the acrAB-TolC pump was brought to over-expression of the efflux gene, by adapting it gradually to higher concentrations of enrofloxacin, a commonly used veterinary antibiotic, until it was able to survive thousand times the initial concentration (0.06 to 60 mg/L).
This Salmonella strain overexpresses the efflux gene,making it resistant to a wide range of antibiotics (e.g.tetracyclines, ß-lactams). For substance screening, the strain was stained with a fluorescent dye in the presence of potential EPIs.
After adding glucose, which induces efflux activity,the shifts in fluorescence were measured (Figure 3). In the untreated control, the dye was extruded and fluorescence rapidly decreased as a result. When Salmonella is treated with known EPIs, phenylalanine-arginine ß-naphthylamide (PAßN) and the anti-malaria drug artesunate, the efflux is clearly inhibited.
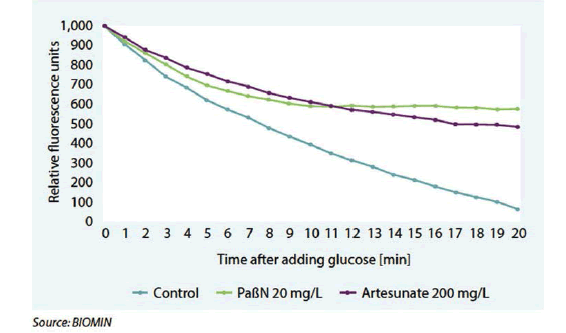
Gut resistome studies to assessantibiotic resistance on the farm
The gastrointestinal tract is habitat of an enormous species diversity and density, a reservoir for thousands of antibiotic resistance genes. Assessment of antibiotic resistance has long relied on traditional isolation techniques by cultivating and counting bacteria on nutrient agars with and without antibiotics.
However, these provide only information on the minority of bacteria—those that can grow under laboratory conditions.
Quantitative polymerase chain reaction
Molecular methods, targeting the genetic basis of resistance,use DNA to characterize and quantify antibiotic resistance determinants. Extraction of DNA from farm-derived samples (e.g. feces) allows reliable quantification of resistance genes in a high number of samples,by using quantitative Polymerase Chain Reaction (qPCR) (Figure 4). PCR is a targeted approach using synthetic oligonucleotides ("primers") that are complementary to the flanking regions of the gene of interest to amplify this particular gene fragment. It will not provide information on the presence of genes that are not targeted by the primers.
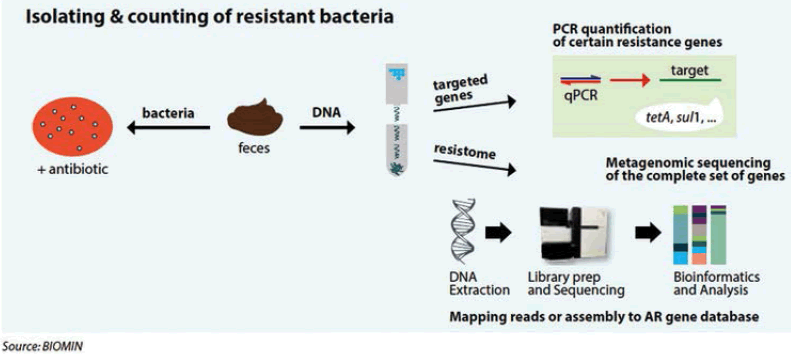
Metagenomics offers a complete view
The most comprehensive approach to exploring antibiotic resistance in complex environments uses metagenomics, which aims to assess the entire genomic information stored in a given sample ("metagenome") by using modern sequencing technologies. Current platforms, like the Illumina HiSeq, yield anywhere from 10 to 1000 GB of DNA sequences in a single lane. The sequencing datasets can be analyzed by assembly of the short reads into larger contiguous DNA fragments orby mapping to reference sequences. This method allows determination of the microbiota composition and simultaneous detection and quantification of the complete set of resistance genes ("resistome") or other genes of interest (e.g. virulence) in the microbiota.
For more information, please visit: http://www.biomin.net/en/magazines/science-solutions-special-issue-rd/
For more of the article, please click here.